Pfu DNA Polymerase, cloned
Catalog #: 091316-500U
Size: 500 units
Concentration: 2.5 U/ml
Store at -20°C
For Research use only
DESCRIPTION
Pfu DNA Polymerase is a highly accurate, thermostable DNA polymerase with 3'→5' exonuclease activity. This activity allows the enzyme to correct errors (proofreading) during DNA amplification, resulting in significantly higher fidelity compared to polymerases lacking this function. Pfu DNA polymerase does not have a 5'→3' exonuclease activity, and lacks a terminal transferase activity (or a non-templated nucleotide addition activity).
Applications
- High-fidelity PCR
- Molecular cloning
- RT-PCR for cDNA cloning
- Primer extension
- Site-directed mutagenesis.
Source
Pyrococcus furiosus
Storage Buffer
This enzyme is supplied in a buffer containing 20 mM Tris-HCl (pH 8.2), 100 mM KCl, 1 mM DTT, 0.1 mM EDTA, 0.1% Triton X-100, 0.1% Tween-20, and 50% glycerol.
Storage Condition
Store Pfu DNA Polymerase at -20°C. Avoid repeated freeze-thaw cycles.
10X Cloned Pfu Buffer
200 mM Tris-HCl (pH 8.8 at 25°C), 100 mM KCl, 100 mM (NH4)2SO4, 20 mM MgSO4, 1% Triton X-100, and 1 mg/ml BSA.
Unit Definition
One unit is defined as the amount of enzyme that incorporates 10 nmol of dNTPs in 30 minutes at 72°C under standard assay conditions. Please see below for details.
PCR PROTOCOL
This is a general protocol and may need to be optimized for your specific application.
Materials:
Materials provided by the kit:
- Pfu DNA Polymerase (2.5 U/µL)
- 10x Pfu Reaction Buffer (containing 2 mM MgSO4)
- 50 mM MgSO4
- 100% DMSO
Materials provided by the user:
- dNTP Mix (e.g., 2 mM of each dNTP)
- Forward Primer (e.g., 10 µM)
- Reverse Primer (e.g., 10 µM)
- Template DNA
- Nuclease-free water
Procedure:
1. Prepare the reaction mixture on ice (50 µL total):
|
Component |
Volume (per reaction) |
Final Concentration |
|
10x Pfu Buffer |
5 µL |
1x |
|
dNTP Mix (2 mM) |
5 µL |
200 µM each |
|
Forward Primer |
3 µL |
0.6 µM |
|
Reverse Primer |
3 µL |
0.6 µM |
|
Template DNA |
0.1-100 ng |
Variable |
|
Pfu DNA Polymerase |
1 µL |
2.5 U/µL |
|
Nuclease-free water |
Add to a volume of 50 µL |
2. Mix gently by pipetting multiple times.
3. Perform PCR using the following cycling parameters:
|
Step |
Temperature (°C) |
Time |
Cycles |
|
Initial Denaturation |
94 |
2 min |
1 |
|
Denaturation |
94 |
30 sec |
25-30 |
|
Annealing |
45-65 |
30 sec |
|
|
Extension |
72 |
1 min/kb |
|
|
Final Extension |
72 |
10 min |
1 |
|
Hold |
4-15 |
Vary |
1 |
- Annealing Temperature: Optimize based on primer Tm. Start 5°C below the lower Tm of the two primers.
- Extension Time: Approximately 1 minute per kilobase of the expected PCR product.
- Number of Cycles: 25-30 cycles are usually sufficient.
4. Analyze the PCR product by agarose gel electrophoresis.
Optimization:
- Magnesium Concentration: Optimal MgSO4 concentration is crucial. Varying it between 2 mM and 4 mM may be necessary. The supplied 10X Pfu reaction buffer contains 2 mM MgSO4, but additional MgSO4 may be required for optimal reaction efficiency.
- Annealing Temperature: Critical for PCR amplification. If a standard reaction condition does not work well for a specific PCR amplification, experiments may need to be performed to determine the optimal annealing temperature.
- Primer Design: Use high-quality primers with a GC content of 40-60% and minimal secondary structure.
- Template Quality: Use high-quality, purified DNA template.
- Enzyme Concentration: Adjust enzyme concentration if necessary.
- Additives: A vial of DMSO is provided, which may improve amplification of DNA template containing a high GC content or strong secondary structure.
Troubleshooting:
- No PCR product: Check all reagents, optimize annealing temperature and MgSO4 concentration, ensure template quality.
- Non-specific PCR products: Increase annealing temperature, and/or redesign primers.
- Weak or faint bands: Increase the number of cycles, optimize MgSO4 concentration, ensure template quality, or increase template amount.
QUALITY CONTROL ANALYSIS
Activity determination
Pfu DNA polymerase activity was determined in a 50 µL reaction containing: 20 mM Tris-HCl (pH 8.8 at 25°C), 10 mM KCl, 10 mM (NH4)2SO4, 2 mM MgSO4, 0.1% Triton X-100, and 0.1 mg/ml BSA, 0.2 mM each of dNTPs, a pair of long oligos that anneal with each other at the 3’ ends (16 ng/µl), and 2 µL of enzyme (with serial dilutions). The reaction mixture was incubated at 72°C for 30 minutes. Amount of double-stranded DNA (dsDNA) generated by Pfu polymerase is measured using a fluorometer in the presence of a dsDNA-specific dye. One unit of the enzyme is defined as the amount that incorporates 10 nmol of dNTPs in 30 minutes at 72°C.
Functional Assay
Pfu DNA polymerase was used for PCR reactions in a 50 µL volume containing: 20 mM Tris-HCl (pH 8.8 at 25°C), 10 mM KCl, 10 mM (NH4)2SO4, 2 mM MgSO4, 0.1% Triton X-100, and 0.1 mg/ml BSA, 0.2 mM each of dNTPs, 10 ng phage l DNA, a pair of oligos (300 ng each), and 2.5 unit of enzyme. The oligo pairs anneal to the l DNA and the expected products are 300, 500, 700, 1000, and 1500 bp in length, respectively. The PCR reaction was performed under the following conditions: 1X (94°C, 2 min); 25X (94°C, 30 s; 45°C, 30 s; 72°C, 2 min); 1X (72°C, 10 min); 1X (15°C, hold). The PCR products are resolved in an agarose gel containing ethidium bromide and visualized.
Lack of Nuclease Contamination
To test for potential nuclease contamination, 5.0 µg of an EcoRI-linearized plasmid (pSK3000; 3.0 kb) was incubated with 2.5 unit of Pfu DNA polymerase in a 50 µL reaction containing 20 mM Tris-HCl (pH 8.8 at 25°C), 10 mM KCl, 10 mM (NH4)2SO4, 2 mM MgSO4, 0.1% Triton X-100, and 0.1 mg/ml BSA, 0.2 mM each of dNTPs. The reaction was first incubated at 37°C for 5.5 hr, followed by further incubation at room temperature for 5 days. An aliquot of 10 µL was resolved in a 1% agarose gel containing ethidium bromide. No DNA degradation was detected.
Lack of Detectable DNA Contamination
To test for potential DNA contamination, 2.5 unit of Pfu DNA polymerase was used for PCR reactions in the presence (positive control) or absence of the Pfu expression plasmid. The reactions were carried out in a 50 µL volume containing: 20 mM Tris-HCl (pH 8.8 at 25°C), 10 mM KCl, 10 mM (NH4)2SO4, 2 mM MgSO4, 0.1% Triton X-100, and 0.1 mg/ml BSA, 0.2 mM each of dNTPs, a pair of Pfu-specific oligos (300 ng each), 2.5 unit of enzyme, and ±1 ng of plasmid pDOF2-Pfu. The oligo pair was designed to amplify a 1.7 kb fragment of the Pfu ORF at the 3’ end. The PCR reactions were performed under the following conditions: 1X (94°C, 2 min); 30X (94°C, 30 s; 50°C, 30 s; 72°C, 2 min); 1X (72°C, 10 min); 1X (15°C, hold). The reaction products were resolved in an agarose gel containing ethidium bromide and visualized. PCR product was detected only in the reaction containing the template plasmid (positive control), suggesting that there was no detectable DNA contamination in 2.5 unit of Pfu.
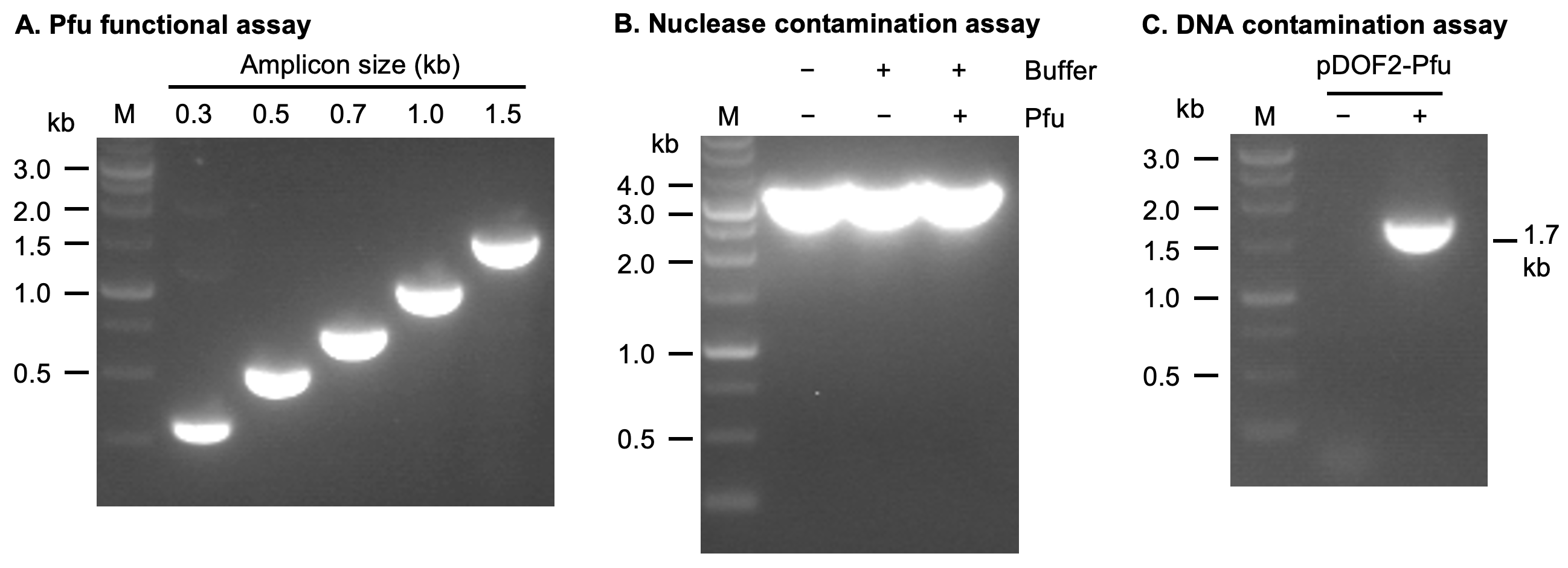
Figure 1. Quality control assays of the Pfu DNA polymerase. (A) Functional assay. 2.5 U of Pfu DNA polymerase was used to amplify DNA fragments from phage lambda DNA. The amplicon sizes are 0.3, 0.5, 0.7, 1.0, and 1.5 kb, respectively. (B) Nuclease contamination assay. A 3.0 kb linearized plasmid was incubated with and without Pfu DNA polymerase for prolonged time. No DNA degradation was observed. (C) DNA contamination assay. PCR reactions were performed with 2.5 U of Pfu DNA polymerase in the presence of Pfu gene-specific primers. PCR product was observed only when the expression plasmid (pDOF2-Pfu) was added as the template (positive control), suggesting there is no detectable DNA contamination in the enzyme.
PRODUCT USE LIMITATION and DISCLAIMER
This product is developed for in vitro research use only. It has not been tested for diagnostic purposes or other applications. The performance of this enzyme is guaranteed for its intended use only. All reagents provided should be handled according to established laboratory safety procedures.
The purchase of this product gives the buyer the right to use the reagents internally. Buyer agrees not to transfer the product to third parties (such as resell or repackage).
This instruction is for informational purposes only and is subject to change without notice.



